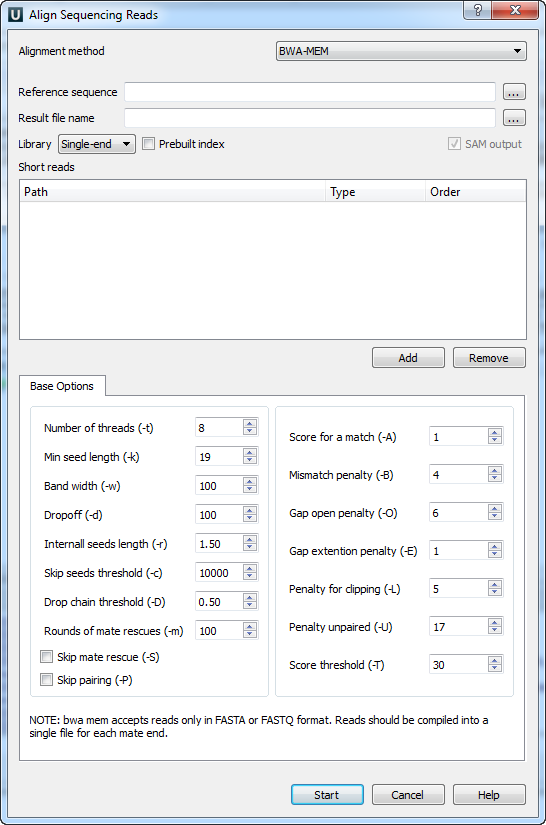
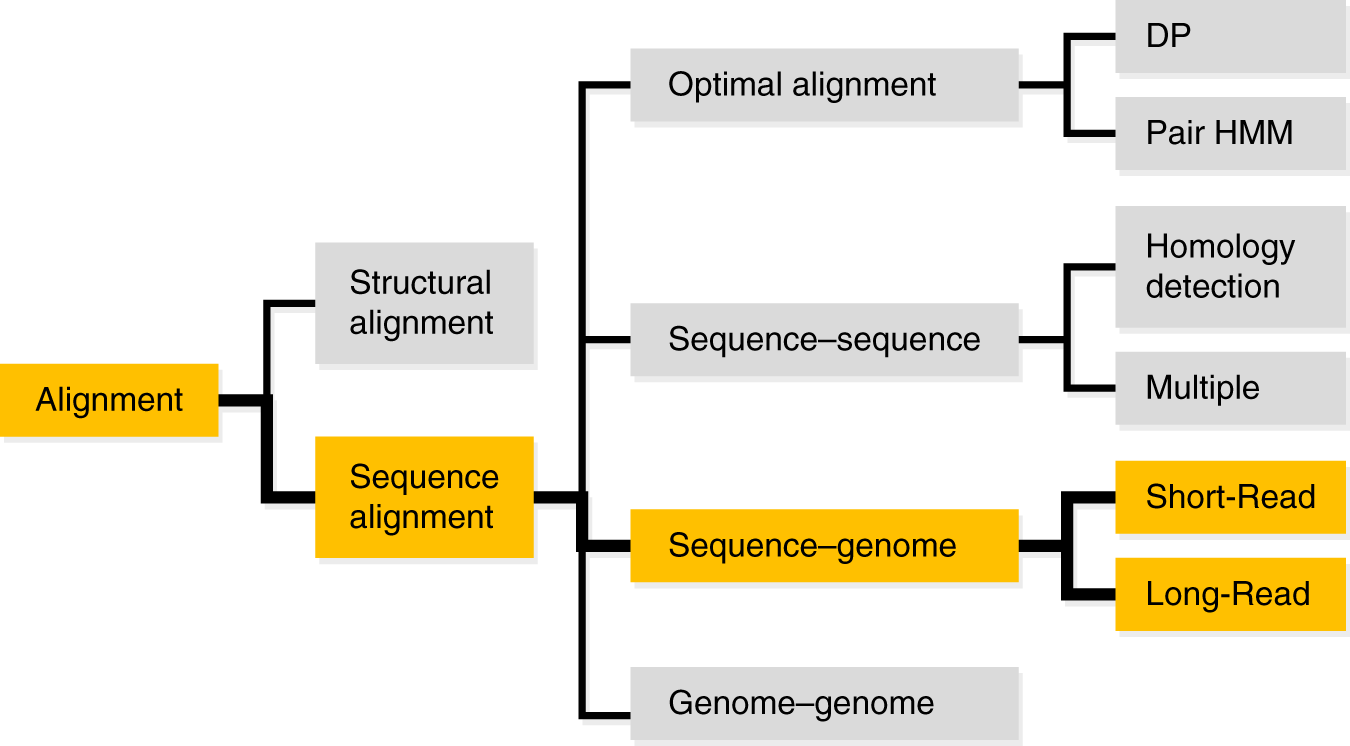
myExperiment - Workflows - cutadapt+BWA-MEM-mis1-i2,2-gapex2,2+filter_c5.0_c4.15+idxStats (Marc Ciosi) [Galaxy Workflow]
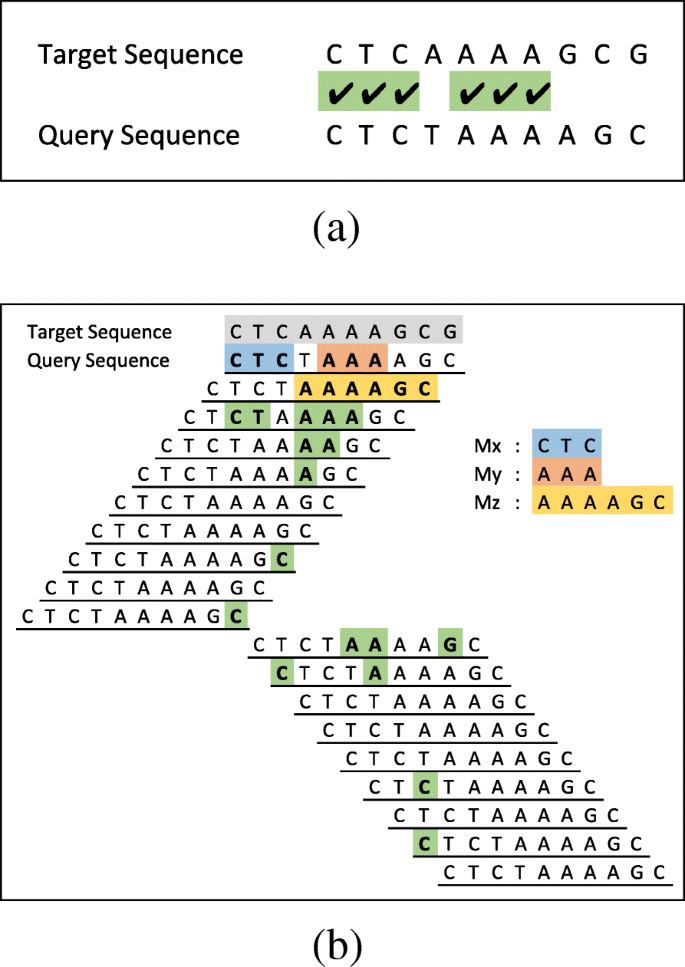
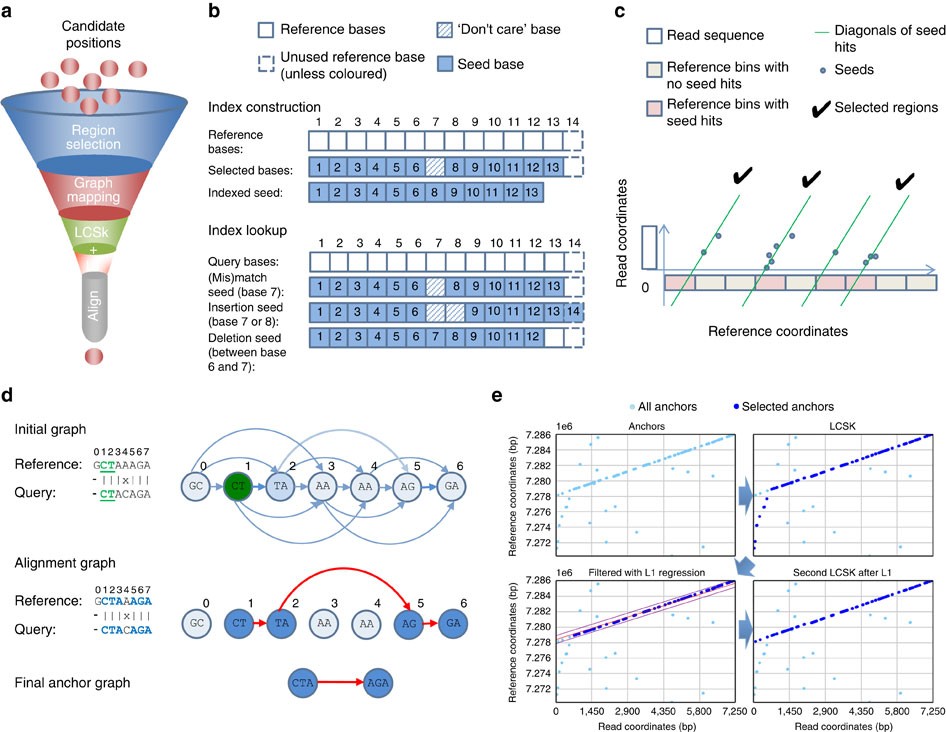
Pairwise alignment of nucleotide sequences using maximal exact matches | BMC Bioinformatics | Full Text

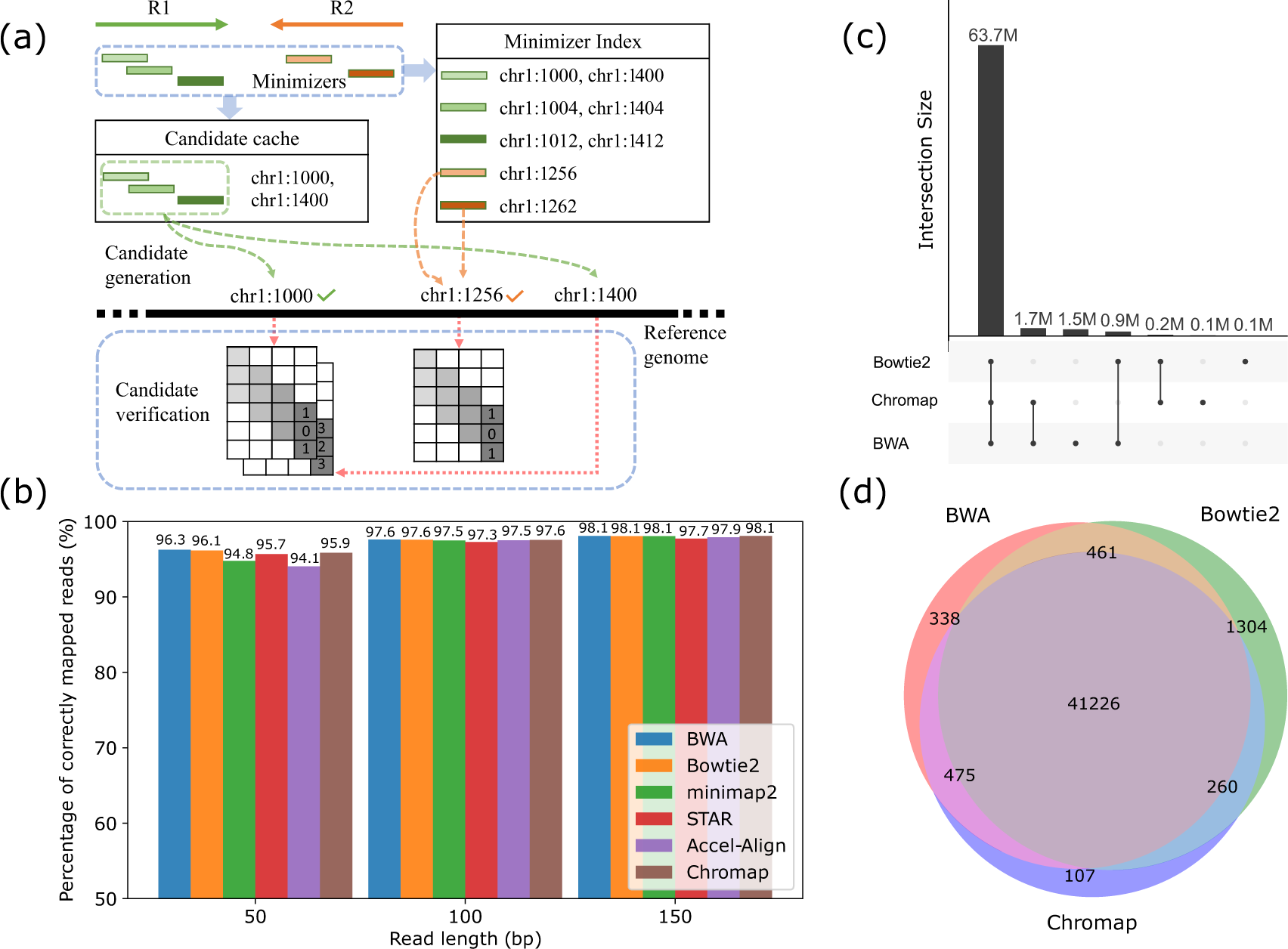
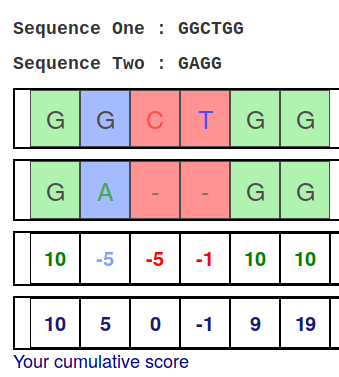
An example SW score matrix is shown (penalties for match, mismatch, gap... | Download Scientific Diagram